Or fancy words that mean very simple things.
At the heart of most data mining, we are trying to represent complex things in a simple way. The simpler you can explain the phenomenon, the better you understand. It’s a little zen – compression is the same as understanding.
Warning: Some math ahead.. but stick with it, it’s worth it.
When faced with a matrix of very large number of users and items, we look to some classical ways to explain it. One favorite technique is Singular Value Decomposition, affectionately nicknamed SVD. This says that we can explain any matrix in the form
Here
is of size num_users and num_products
is of size num_users and num_users
is a rectangular diagonal matrix of size num_users and num_products
is of size num_products and num_products
This is cool because the numbers in the diagonal are decreasing values of variance (roughly speaking). So the first number captures the most variance in
, the second, less so, and so on, till all the numbers put together capture all the variance of
.
You see where this is going. If we are comfortable with explaining only 10% of the variance, we can do so by only taking some of these values. We can then compress .
The other nice thing about all this is eigenvectors (which those roughly represent) are orthogonal. In other words, they are perpendicular to each other and there aren’t any mixed effects. If you don’t understand this paragraph, don’t worry. Keep on.
Consider that above can be rewritten as
or –
If is of size num_users and num_categories and
is of size num_products and num_categories, then we have a decent model that approximates
into
.
is now called a low rank approximation of
, or truncated SVD.
It is important to understand what these matrices represent. is the representation of users in some low dimension space (say romance, action). And
is the representation of products in the very same low dimension space. However, we don’t know exactly what these factors mean. They could be romance/action or they could NYC/Dallas. This is also why this method is sometimes called Latent Factor Matrix Factorization.. wow, quite a mouthful.
In my chapter in the book Data Mining Applications with R, I go over different themes of matrix factorization models (and other animals as well). But for now, I am only going to cover the basic one that works very well in practice. And yes, won the Netflix prize.
There is one problem with our formulation – SVD is only defined for dense matrices. And our matrix is usually sparse.. very sparse. Netflix’s challenge matrix was 1% dense, or 99% sparse. In my job at Rent the Runway, it is only 2% dense. So what will happen to our beautiful formula?
Machine learning is sort of a bastard science. We steal from beautiful formulations, complex biology, probability, Hoefding’s inequality, and derive rules of thumb from it. Or as an artist would say – get inspiration.
So here is what we are going to do. We are going to ignore all the null values when we solve this model. Our cost function now becomes
Here is the difference between observed data and our prediction.
is simply a matrix with 1 where we have a value and 0 where we do not. Multiplying these two we are only considering the cost when we observe a value. We are using L2 regularization of magnitude
. We are going to divide by 2 to make all other math easier. The cost is relative so it doesn’t matter.
Using this our gradients become –
If we were wanted to minimize the cost function, we can move in the direction opposite to the gradient at each step, getting new estimates for and
each time.
This looks easy enough. One last thing. We now have what we want to minimize, but how do we do it? R provides many optimization tools. There is a whole CRAN page on it. For our purpose we will use out of the box optim() function. This allows us to access a fast optimization method L-BFGS-B. It’s not only fast, but also doesn’t take too memory intensive which is desirable in our case.
We need to give it the cost function and the gradient that we have above. It also expects one gradient function, so we need to unroll it out to do both our gradients.
This file contains hidden or bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| unroll_Vecs <- function (params, Y, R, num_users, num_movies, num_features) { | |
| # Unrolls vector into X and Theta | |
| # Also calculates difference between preduction and actual | |
| endIdx <- num_movies * num_features | |
| X <- matrix(params[1:endIdx], nrow = num_movies, ncol = num_features) | |
| Theta <- matrix(params[(endIdx + 1): (endIdx + (num_users * num_features))], | |
| nrow = num_users, ncol = num_features) | |
| Y_dash <- (((X %*% t(Theta)) – Y) * R) # Prediction error | |
| return(list(X = X, Theta = Theta, Y_dash = Y_dash)) | |
| } | |
| J_cost <- function(params, Y, R, num_users, num_movies, num_features, lambda, alpha) { | |
| # Calculates the cost | |
| unrolled <- unroll_Vecs(params, Y, R, num_users, num_movies, num_features) | |
| X <- unrolled$X | |
| Theta <- unrolled$Theta | |
| Y_dash <- unrolled$Y_dash | |
| J <- .5 * sum( Y_dash ^2) + lambda/2 * sum(Theta^2) + lambda/2 * sum(X^2) | |
| return (J) | |
| } | |
| grr <- function(params, Y, R, num_users, num_movies, num_features, lambda, alpha) { | |
| # Calculates the gradient step | |
| # Here lambda is the regularization parameter | |
| # Alpha is the step size | |
| unrolled <- unroll_Vecs(params, Y, R, num_users, num_movies, num_features) | |
| X <- unrolled$X | |
| Theta <- unrolled$Theta | |
| Y_dash <- unrolled$Y_dash | |
| X_grad <- (( Y_dash %*% Theta) + lambda * X ) | |
| Theta_grad <- (( t(Y_dash) %*% X) + lambda * Theta ) | |
| grad = c(X_grad, Theta_grad) | |
| return(grad) | |
| } | |
| # Now that everything is set up, call optim | |
| print( | |
| res <- optim(par = c(runif(num_users * num_features), runif(num_movies * num_features)), # Random starting parameters | |
| fn = J_cost, gr = grr, | |
| Y=Y, R=R, | |
| num_users=num_users, num_movies=num_movies,num_features=num_features, | |
| lambda=lambda, alpha = alpha, | |
| method = "L-BFGS-B", control=list(maxit=maxit, trace=1)) | |
| ) |
This is great, we can iterate a few times to approximate users and items into some small number of categories, then predict using that.
I have coded this into another package recommenderlabrats.
Let’s see how this does in practice against what we already have. I am going to use the same scheme as last post to evaluate these predictions against some general ones. I am not using Item Based Collaborative Filtering because it is very slow
This file contains hidden or bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| require(recommenderlab) # Install this if you don't have it already | |
| require(devtools) # Install this if you don't have this already | |
| # Get additional recommendation algorithms | |
| install_github("sanealytics", "recommenderlabrats") | |
| data(MovieLense) # Get data | |
| # Divvy it up | |
| scheme <- evaluationScheme(MovieLense, method = "split", train = .9, | |
| k = 1, given = 10, goodRating = 4) | |
| scheme | |
| # register recommender | |
| recommenderRegistry$set_entry( | |
| method="RSVD", dataType = "realRatingMatrix", fun=REAL_RSVD, | |
| description="Recommender based on Low Rank Matrix Factorization (real data).") | |
| # Some algorithms to test against | |
| algorithms <- list( | |
| "random items" = list(name="RANDOM", param=list(normalize = "Z-score")), | |
| "popular items" = list(name="POPULAR", param=list(normalize = "Z-score")), | |
| "user-based CF" = list(name="UBCF", param=list(normalize = "Z-score", | |
| method="Cosine", | |
| nn=50, minRating=3)), | |
| "Matrix Factorization" = list(name="RSVD", param=list(categories = 10, | |
| lambda = 10, | |
| maxit = 100)) | |
| ) | |
| # run algorithms, predict next n movies | |
| results <- evaluate(scheme, algorithms, n=c(1, 3, 5, 10, 15, 20)) | |
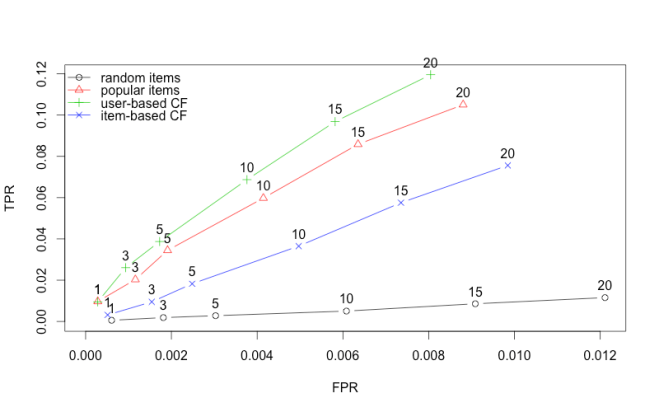
| # Draw ROC curve | |
| plot(results, annotate = 1:4, legend="topleft") | |
| # See precision / recall | |
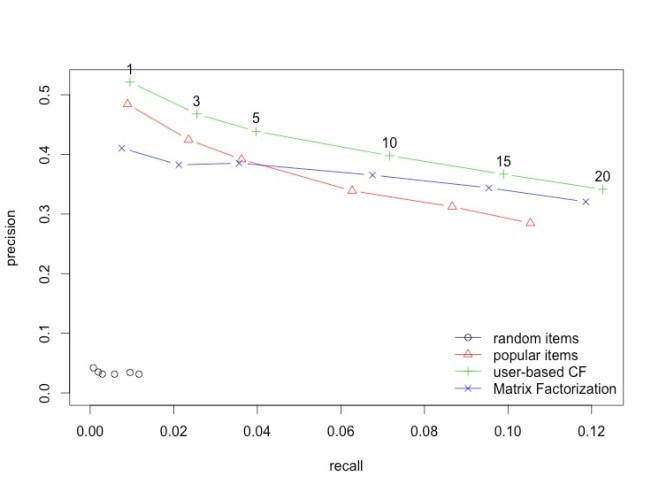
| plot(results, "prec/rec", annotate=3) |
It does pretty well. It does better than POPULAR and is equivalent to UBCF. So why use this over UBCF or the other way round?
This is where things get interesting. This algorithm can be sped up quite a lot and more importantly, parallelised. It uses way less memory than UBCF and is more scalable.
Also, if you have already calculated , i.e. your items in some lower dimensional space, you might get away with just putting the users in that space. Now things become really fast because all you have to do is keep
fixed and figure out
.
I have gone ahead and implemented a version where we just calculate , I leave it to you as an exercise to modify the code above to test this out as well. The algorithm is called RSVD_SPLIT. Also feel free to try other values of categories, lambda and maxit and see how things change.
On the other hand, the latent categories are very hard to explain. For UBCF you can say this user is similar to these other users. For IBCF, one can say this item that the user picked is similar to these other items. But that not the case for this particular flavor of matrix factorization. We will re-evaluate these limitations later.
The hardest part for a data scientist is to disassociate themselves from their dear models and watch them objectively in the wild. Our simulations and metrics are always imperfect but necessary to optimize. You might see your favorite model crash and burn. And a lot of times simple linear regression will be king. The job is to objectively measure them, tune them and see which one performs better in your scenario.
Good luck.